Fast PFNETs in Erlang
Pathfinder Networks - and how to implement them in Erlang
February 14, 2009
programming
erlang
Introduction
Pathfinder Networks (PFNETs) are networks derived from a graph representing proximity data. Basically, each node is connected to (almost) every other node by a weighted link, and that makes it hard to see what’s going on. The Pathfinder algorithm prunes the graph by removing links which are weighted higher than another path between the same nodes.
For example: A links to B with weight 5. A also links to C with weight 2, and C links to B with weight 2. Adding up the weights, A to B direct is 5, A to C to B is 4. Result: we remove the link from A to B, as the route via C is shorter. There are different ways to calculate the path lengths (using the Minkowski r-metric), but you get the general idea. The resulting PFNET has fewer links and is easier to analyse.
In Schvaneveldt (1990) an algorithm for computing PFNETs is given, but it is rather complex and computationally intensive. There is an improved algorithm called Binary Pathfinder, but that is apparently more memory intensive. Not very promising so far, but then along comes A new variant of the Pathfinder algorithm to generate large visual science maps in cubic time, by Quirin, Cordón, Santamaría, Vargas-Quesada, and Moya-Anegón. This algorithm is a lot faster (by about 450 times), but has one disadvantage: speed is traded in for flexibility. The original Pathfinder algorithm has two parameters, r (the value of the Minkowski metric to be used) and q (the maximum length of paths to be considered). The fast algorithm only has r, and always uses the maximum value for q (which is n-1). I know too little about the application of PFNETs to say whether this is important at all; for the uses I can envisage it does not seem to matter.
As added bonus, the algorithm in pseudo code in Quirin et al. is very short and straightforward. They’re using a different shortest-path algorithm to identify, erm, shorter paths. And then it’s very simple to prune the original network.
A picture tells more than 1K words, so here instead of 2000 words the before and after, graph layout courtesy of the graphviz program:
A network example (from Schvaneveldt 1990)
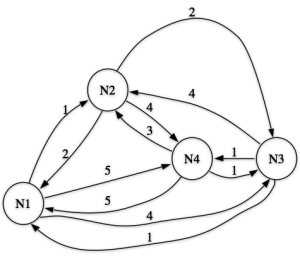 Fig 1: A network example (from Schvaneveldt 1990)
Fig 1: A network example (from Schvaneveldt 1990)
Here, each node is linked to every other node. Running the PFNET algorithm on it, we get the output shown in the second figure.
A PFNET generated from the previous graph:
 Fig 2: A PFNET generated from the previous graph
Fig 2: A PFNET generated from the previous graph
If you compare the output with the actual result from Schvaneveldt’s book (p.6 / 7), you’ll realise that it is not identical, and the reason for that is that the example there limits the path-length, using the parameters (r = 1, q = 2) rather than (r = 1, q = n-1) as in the example shown here. As a consequence, the link from N1 to N4 (with a weight of 5) disappears, because of the shorter path (N1-N2-N3-N4, weight 4). But that path is too long if q is just 2, and so it is kept in Schvaneveldt’s example.
Implementation
It is not possible to implement the pseudo-code given in Quirin et al directly (in Erlang), as they use destructive updates of the link matrix, which we obviously cannot do in Erlang. But the first, naïve, implementation is still quite short. The input graph is represented as a matrix (n x n) where each value stands for the link weight, with zero being used to indicate non-existing links. I have written a function that creates a dot file from a matrix, which is then fed into graphviz for generating images as the ones shown above.
There are basically two steps: creating a matrix of shortest paths from the input matrix, and then generating the PFNET by comparing the two matrices; if a certain cell has the same value in both matrices, then it is a shortest path and is kept, otherwise there is a shorter path and it’s pruned. Here is the main function:
find_path(Matrix) ->
Shortest = loop1(1,length(Matrix),Matrix),
generate_pfnet(Matrix, Shortest, []).
Next we have the three loops (hence ‘cubic time’!) of the Floyd-Warshall shortest path algorithm to create the shortest path matrix:
loop1(K,N,Matrix) when K > N ->
Matrix;
loop1(K,N,Matrix) ->
NewMatrix = loop2(K,1,Matrix,
Matrix,[]),
loop1(K+1,N,NewMatrix).
loop2(_,_,_,[],Acc) ->
lists:reverse(Acc);
loop2(K,I,D,[Row|Rest],Acc) ->
NewRow = loop3(K,I,1,D, Row, []),
loop2(K,I+1,D,Rest,[NewRow|Acc]).
loop3(_,_,_,_, [], Acc) ->
lists:reverse(Acc);
loop3(K,I,J,D, [X|Rest], Acc) ->
loop3(K,I,J+1,D, Rest,
[min(X, get(I,K,D) + get(K,J,D))|Acc]).
The final line implements the Minkowski metric with r = 1; this could be expanded to include other values as well, eg r = 2 for Euclidean, or r = ∞ (which seem to be the most common values in use; the latter means using the maximum weight of any path component along the full path).
And here are two utility methods, one to find the smaller of two values, and one to retrieve an element from a matrix (which is a list of rows). There is something of a hack to deal with the fact that zero does not mean a very small weight, but refers to a non-existing link:
min(X, Y) when X < Y -> X;
min(_X, Y) -> Y.
get(Row, Col, Matrix) ->
case lists:nth(Col,
lists:nth(Row, Matrix)) of
0 -> 999999999;
X -> X
end.
And finally, generating the PFNET by comparing the two matrices (again, awful hack included):
generate_pfnet([],[],Result) ->
lists:reverse(Result);
generate_pfnet([R1|Rest1], [R2|Rest2], Acc) ->
Row = generate_pfrow(R1,R2,[]),
generate_pfnet(Rest1, Rest2, [Row|Acc]).
generate_pfrow([],[],Result) ->
lists:reverse(Result);
generate_pfrow([C|Rest1], [C|Rest2], Acc) ->
case C of
999999999 -> C1 = 0;
_ -> C1 = C
end,
generate_pfrow(Rest1, Rest2, [C1|Acc]);
generate_pfrow([C1|Rest1], [C2|Rest2], Acc) ->
generate_pfrow(Rest1, Rest2, [0|Acc]).
Discussion
So this is the basic code. It works, but there is scope for improvement.
- it only currently generates PFNETs with (r = 1, q = n-1)
- there is no parallelism, hence it’s not really making use of Erlang’s strengths.
- the three loops don’t look very elegant, and could probably be replaced by list comprehensions
Because of the matrix being updated, it doesn’t look that easy to
parallelise the processing, but it would work at the level of
updating the individual rows. If that can be done in parallel, it
would probably provide some speed-up (provided the matrix is not
of a trivial size). So the first step would be to change the matrix
processing using lists:map/2, and replacing this then by pmap.
Once the code is up to scratch with full parallelism, and tested on larger matrices I will probably put it up on Google Code in case other people are interested in using it. If you have any suggestions, tell me in the comments!
References
- Schvaneveldt, R. W. (Ed.) (1990)
- Pathfinder Associative Networks: Studies in Knowledge Organization. Norwood, NJ: Ablex. The book is out of print. A copy can be downloaded from the Wikipedia page
- Quirin, A; Cordón, O; Santamaría, J; Vargas-Quesada, B; Moya-Anegón, F (2008)
- “A new variant of the Pathfinder algorithm to generate large visual science maps in cubic time”, Information Processing and Management, 44, p.1611-1623.